The Cooperative Computing Lab

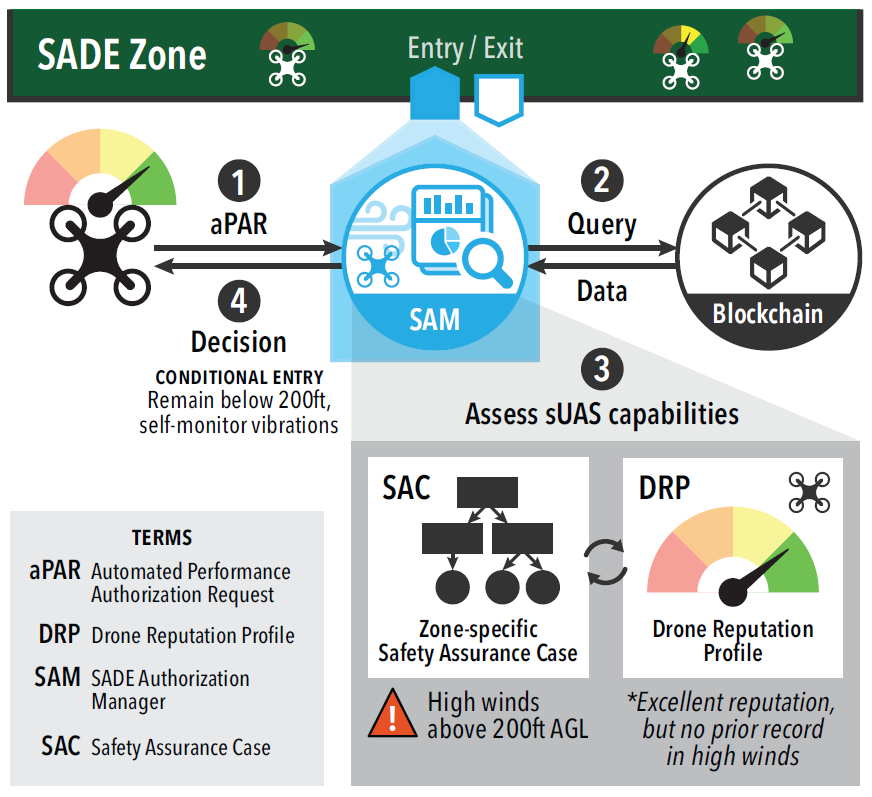
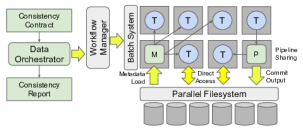
We design software that enables our collaborators to easily harness large scale distributed systems such as clusters, clouds, and grids. We perform fundamental computer science research that enables new discoveries through computing in fields such as physics, chemistry, bioinformatics, biometrics, and data mining.
All of our software is open source and available on GitHub. We are always eager to welcome new collaborators—whether you want to explore our software, contribute to development, or receive support. Explore our tools to see how they can power your distributed computing needs, and visit the getting help section if you need assistance.
CCL Launches Redesigned Website (Jan 06, 2026)
Latest News and Posts
| Apr 16, 2026 | Congratulations to Thanh Son Phung on Passing His Doctoral Defense |
|---|---|
| Apr 09, 2026 | Introducing X-Bucket |
| Mar 31, 2026 | TaskVine Insights: How to Propose a Pull Request to the CCL Team |
| Mar 24, 2026 | Cursor Rules in Research Computing |
| Mar 17, 2026 | TaskVine Insights: A Beginner's Map to the CCTools Codebase and a First Task Dispatch Patch |