The Cooperative Computing Lab

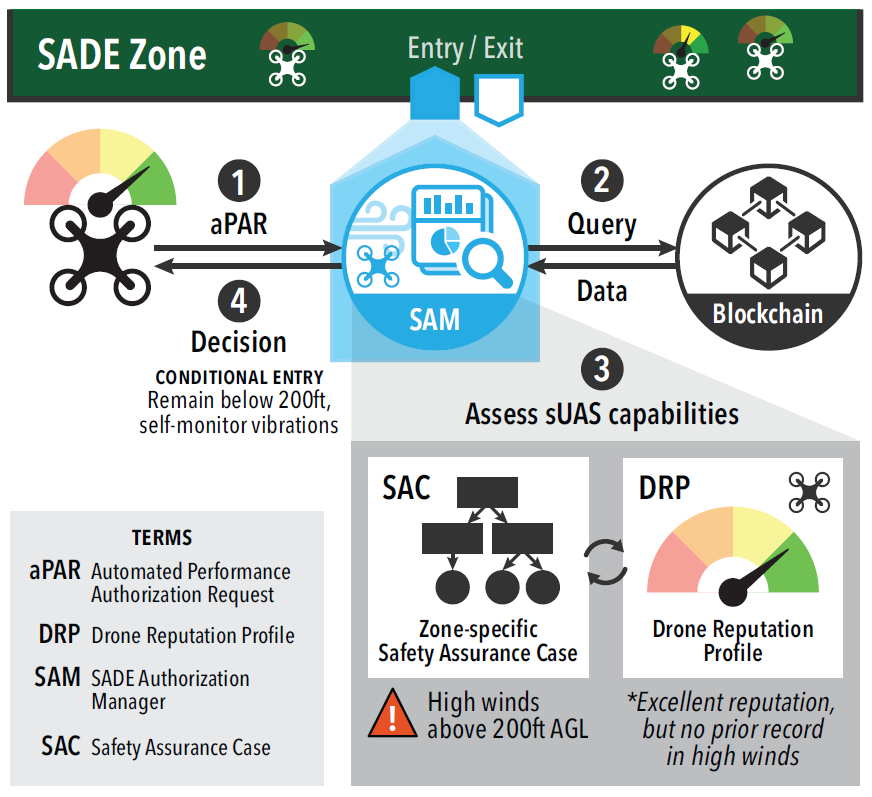
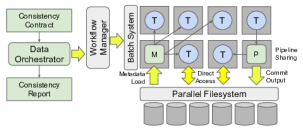
We design software that enables our collaborators to easily harness large scale distributed systems such as clusters, clouds, and grids. We perform fundamental computer science research that enables new discoveries through computing in fields such as physics, chemistry, bioinformatics, biometrics, and data mining.
All of our software is open source and available on GitHub. We are always eager to welcome new collaborators—whether you want to explore our software, contribute to development, or receive support. Explore our tools to see how they can power your distributed computing needs, and visit the getting help section if you need assistance.
CCL Launches Redesigned Website (Jan 06, 2026)
Latest News and Posts
| Jun 03, 2026 | SciWIND at IPDPS 2026 |
|---|---|
| May 12, 2026 | CCL Team at GCASR 2026 |
| May 05, 2026 | TaskVine Insights: Shipping Worker Builds to Remote Nodes and Debugging There |
| Apr 28, 2026 | TaskVine Insights: Submitting Workers to a Cluster |
| Apr 21, 2026 | Congratulations to Colin Thomas on Passing His Candidacy Exam |