
| Patrick Braga-Henebry Computer Science 2009 |
Project: distblast
 distblast is a project created with the intent of harnessing established distributed computing resources in order to parallelize gene sequence matching using the BLAST tool. Gene sequence matching is an I/O- and memory-intensive process and as such cannot be effectively parallelized using only distributed processing resources, i.e. using distributing computing power to parallelize while accessing the same database over the network will result in little performance gain. BLAST is a widely used tool for gene sequence comparisons, and reducing runtime will allow for greater productivity.
distblast is a project created with the intent of harnessing established distributed computing resources in order to parallelize gene sequence matching using the BLAST tool. Gene sequence matching is an I/O- and memory-intensive process and as such cannot be effectively parallelized using only distributed processing resources, i.e. using distributing computing power to parallelize while accessing the same database over the network will result in little performance gain. BLAST is a widely used tool for gene sequence comparisons, and reducing runtime will allow for greater productivity.
BLAST query files are composed of multiple, individual, independent query strings. I composed a suite of shell and Python scripts to divide an incoming query file into multiple query files of roughly equal size for processing via Condor, a distributed platform. The 2.6Gb database was fully replicated maunally onto 16 machines using the Chirp distributed filesystem so as to avoid multiple processes accessing the same database over the network. In order to measure performance increase, query files with 1,2,3..239,240 queries were submitted to distblast. The Unix time command was used to measure total processing time of each query.
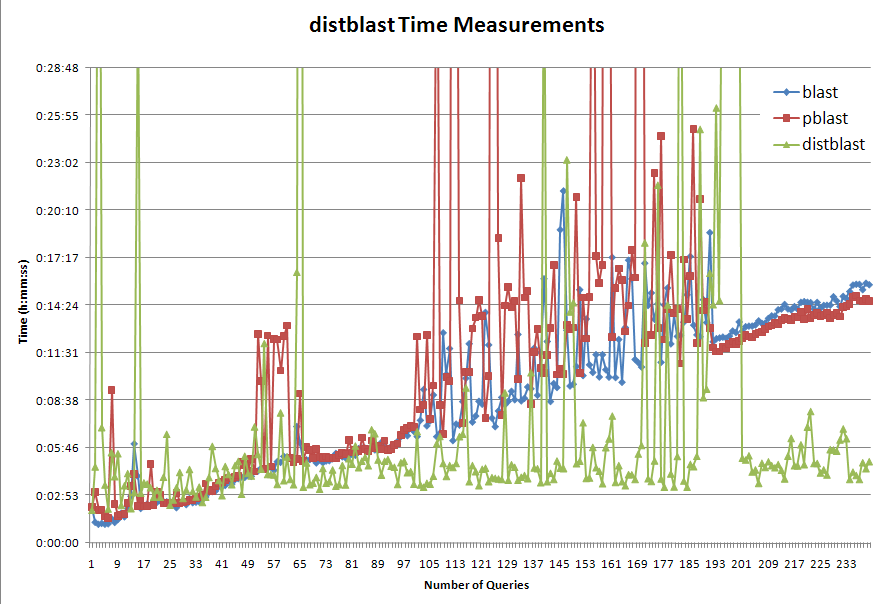
- distblast is the aforementioned set of scripts
- blast is running the n-query job against a single instance of the database residing on the network
- pblast is running a single, undivided n-query job against the locally-stored database
This test shows that distblast does indeed offer performance benefits over using normal blast.
These performance benefits, however, do not yet scale: 240 queries was the limit of this test, and incoming query files were split into batches of 30 queries per job. This results in only 8 machines being utilized at a time. On a 16-machine cluster, the trend for the distblast line should in theory hold until 480 queries before trending upwards because multiple jobs would have to be run on the same machine. Further work includes dynamically replicating the database to multiple machines upon submission of large jobs. In addition, our experiment showed that the number of queries per job could be increased. All the machines used in this project were in the cclsun cluster; in order to scale, the database will have to be reliably replicated to machines outside controlled clusters. Another area for further work is dynamically pulling a fresh, up-to-date copy of the database from its origination source online.